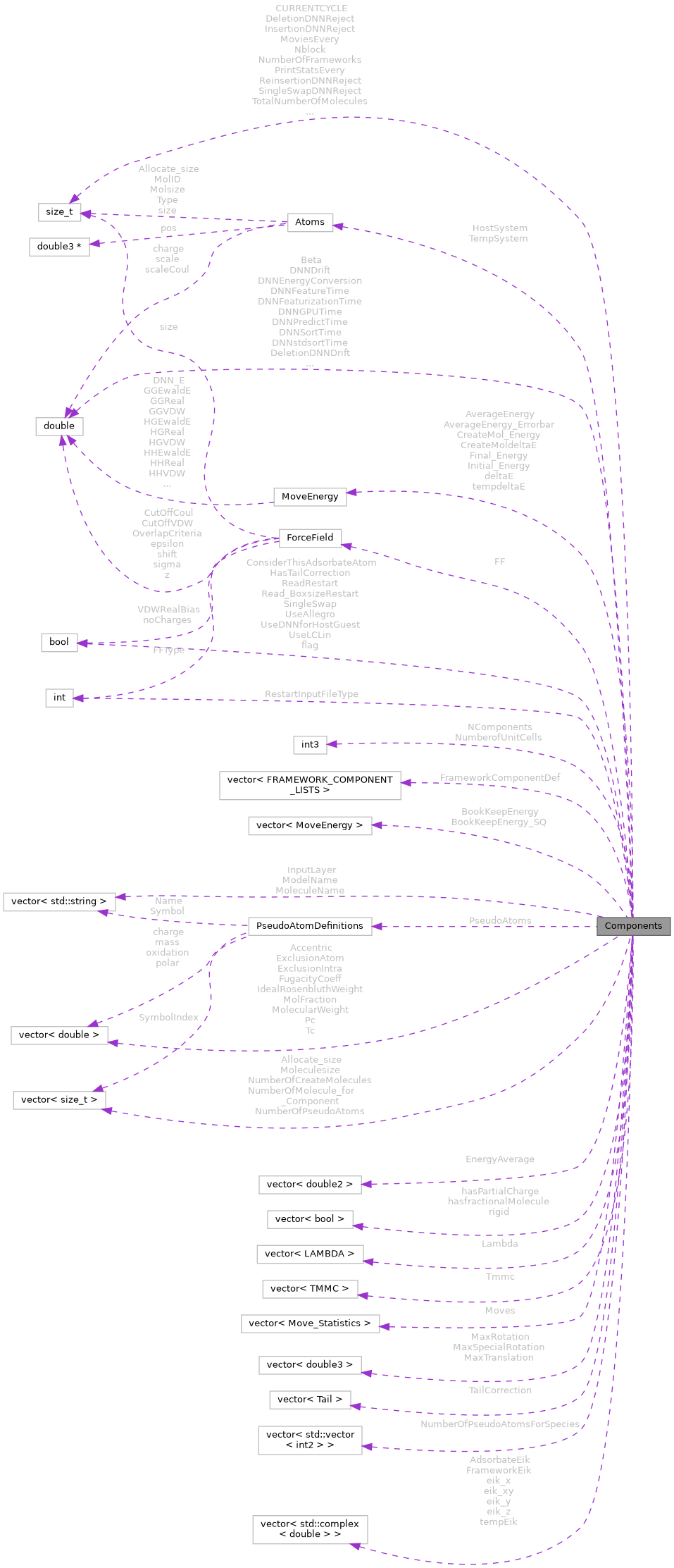
Collaboration diagram for Components:
Public Member Functions | |
| void | Copy_GPU_Data_To_Temp (Atoms GPU_System, size_t start, size_t size) |
| size_t | MatchMoleculeNameToComponentID (std::string Name) |
| void | UpdatePseudoAtoms (int MoveType, size_t component) |
Public Attributes | |
| size_t | Total_Components |
| int3 | NComponents |
| int3 | NumberofUnitCells |
| size_t | MoviesEvery = 5000 |
| size_t | PrintStatsEvery = 5000 |
| size_t | Nblock ={5} |
| size_t | CURRENTCYCLE ={0} |
| double | DNNPredictTime ={0.0} |
| double | DNNFeatureTime ={0.0} |
| double | DNNGPUTime ={0.0} |
| double | DNNSortTime ={0.0} |
| double | DNNstdsortTime ={0.0} |
| double | DNNFeaturizationTime ={0.0} |
| size_t | TotalNumberOfMolecules |
| size_t | NumberOfFrameworks |
| double | Temperature ={0.0} |
| double | Beta |
| bool * | flag |
| std::vector< FRAMEWORK_COMPONENT_LISTS > | FrameworkComponentDef |
| MoveEnergy | Initial_Energy |
| MoveEnergy | CreateMol_Energy |
| MoveEnergy | Final_Energy |
| std::vector< MoveEnergy > | BookKeepEnergy |
| std::vector< MoveEnergy > | BookKeepEnergy_SQ |
| MoveEnergy | AverageEnergy |
| MoveEnergy | AverageEnergy_Errorbar |
| Atoms * | HostSystem |
| Atoms | TempSystem |
| MoveEnergy | tempdeltaE |
| MoveEnergy | CreateMoldeltaE |
| MoveEnergy | deltaE |
| double | FrameworkEwald =0.0 |
| bool | HasTailCorrection = false |
| bool | ReadRestart = false |
| int | RestartInputFileType = RASPA_RESTART |
| bool | Read_BoxsizeRestart = false |
| bool | SingleSwap =false |
| bool | UseDNNforHostGuest = false |
| size_t | TranslationRotationDNNReject =0 |
| size_t | ReinsertionDNNReject =0 |
| size_t | InsertionDNNReject =0 |
| size_t | DeletionDNNReject =0 |
| size_t | SingleSwapDNNReject =0 |
| double | SingleMoveDNNDrift =0.0 |
| double | ReinsertionDNNDrift =0.0 |
| double | InsertionDNNDrift =0.0 |
| double | DeletionDNNDrift =0.0 |
| double | SingleSwapDNNDrift =0.0 |
| double | DNNDrift = 100000.0 |
| double | DNNEnergyConversion |
| bool | UseAllegro = false |
| bool | UseLCLin = false |
| std::vector< std::string > | ModelName |
| std::vector< std::string > | InputLayer |
| size_t * | device_InverseIndexList |
| bool * | ConsiderThisAdsorbateAtom |
| double * | device_Distances |
| std::vector< double2 > | EnergyAverage |
| std::vector< bool > | hasPartialCharge |
| std::vector< bool > | hasfractionalMolecule |
| std::vector< LAMBDA > | Lambda |
| std::vector< TMMC > | Tmmc |
| std::vector< bool > | rigid |
| std::vector< double > | ExclusionIntra |
| std::vector< double > | ExclusionAtom |
| std::vector< std::string > | MoleculeName |
| std::vector< size_t > | Moleculesize |
| std::vector< size_t > | NumberOfMolecule_for_Component |
| std::vector< size_t > | Allocate_size |
| std::vector< size_t > | NumberOfCreateMolecules |
| std::vector< double > | MolFraction |
| std::vector< double > | IdealRosenbluthWeight |
| std::vector< double > | FugacityCoeff |
| std::vector< Move_Statistics > | Moves |
| std::vector< double3 > | MaxTranslation |
| std::vector< double3 > | MaxRotation |
| std::vector< double3 > | MaxSpecialRotation |
| std::vector< double > | Tc |
| std::vector< double > | Pc |
| std::vector< double > | Accentric |
| std::vector< Tail > | TailCorrection |
| std::vector< double > | MolecularWeight |
| ForceField | FF |
| PseudoAtomDefinitions | PseudoAtoms |
| std::vector< size_t > | NumberOfPseudoAtoms |
| std::vector< std::vector< int2 > > | NumberOfPseudoAtomsForSpecies |
| std::vector< std::complex< double > > | eik_xy |
| std::vector< std::complex< double > > | eik_x |
| std::vector< std::complex< double > > | eik_y |
| std::vector< std::complex< double > > | eik_z |
| std::vector< std::complex< double > > | AdsorbateEik |
| std::vector< std::complex< double > > | FrameworkEik |
| std::vector< std::complex< double > > | tempEik |
The documentation for this struct was generated from the following file:
- src_clean/data_struct.h